Plant Chemetics Laboratory
University of Oxford
Welcome to the Plant Chemetics Laboratory
The Plant Chemetics lab investigates the molecular mechanisms underpinning host manipulation by plant pathogens, with a particular focus on apoplast manipulation of Solanaceous plants by bacterial microbes. Our research activities also aim at improved molecular pharming of secreted (glyco)proteins through the depletion of immune responses and by removing secreted proteases. Unique is our approach to display protein activities by pioneering activity-based protein profiling (ABPP), and other chemical proteomic technologies, which offers exciting platforms for collaborations. Please contact us if you are interested to join or collaborate!
News
-
October 2023
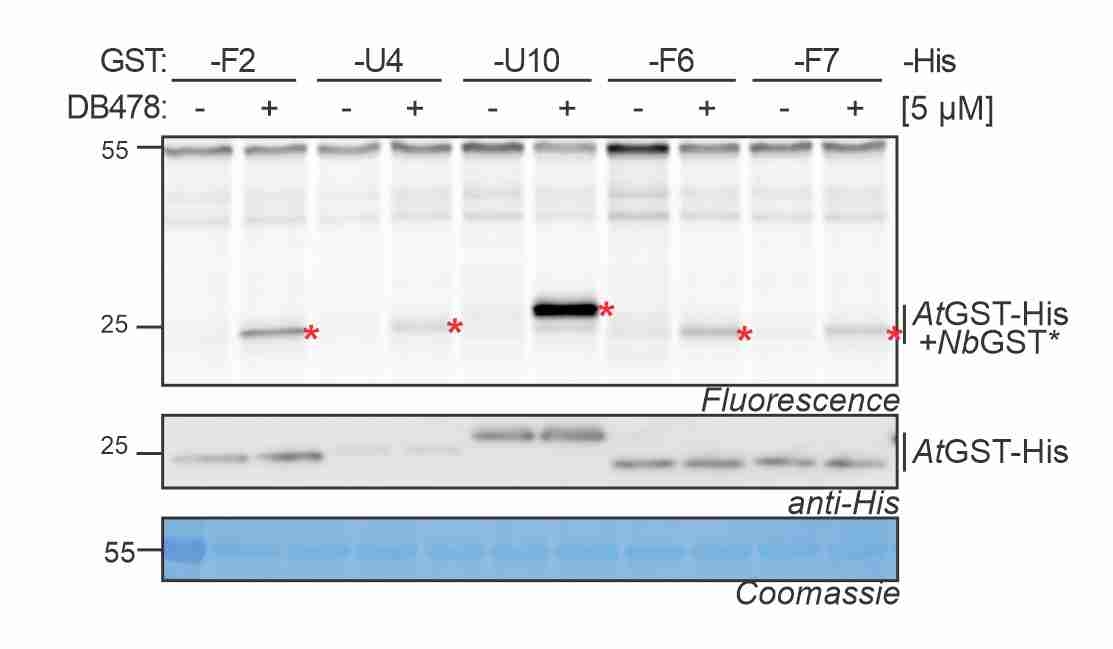
GST photoaffinity labeling! Maria validated a photoaffinity probe for GSTs and studied the GST response to agrochemicals and disease. More here! -
October 2023
NEW BBSRC grant! The BBSRC funds us to continue the validation of novel AI-predicted P68B inhibitors! -
September 2023
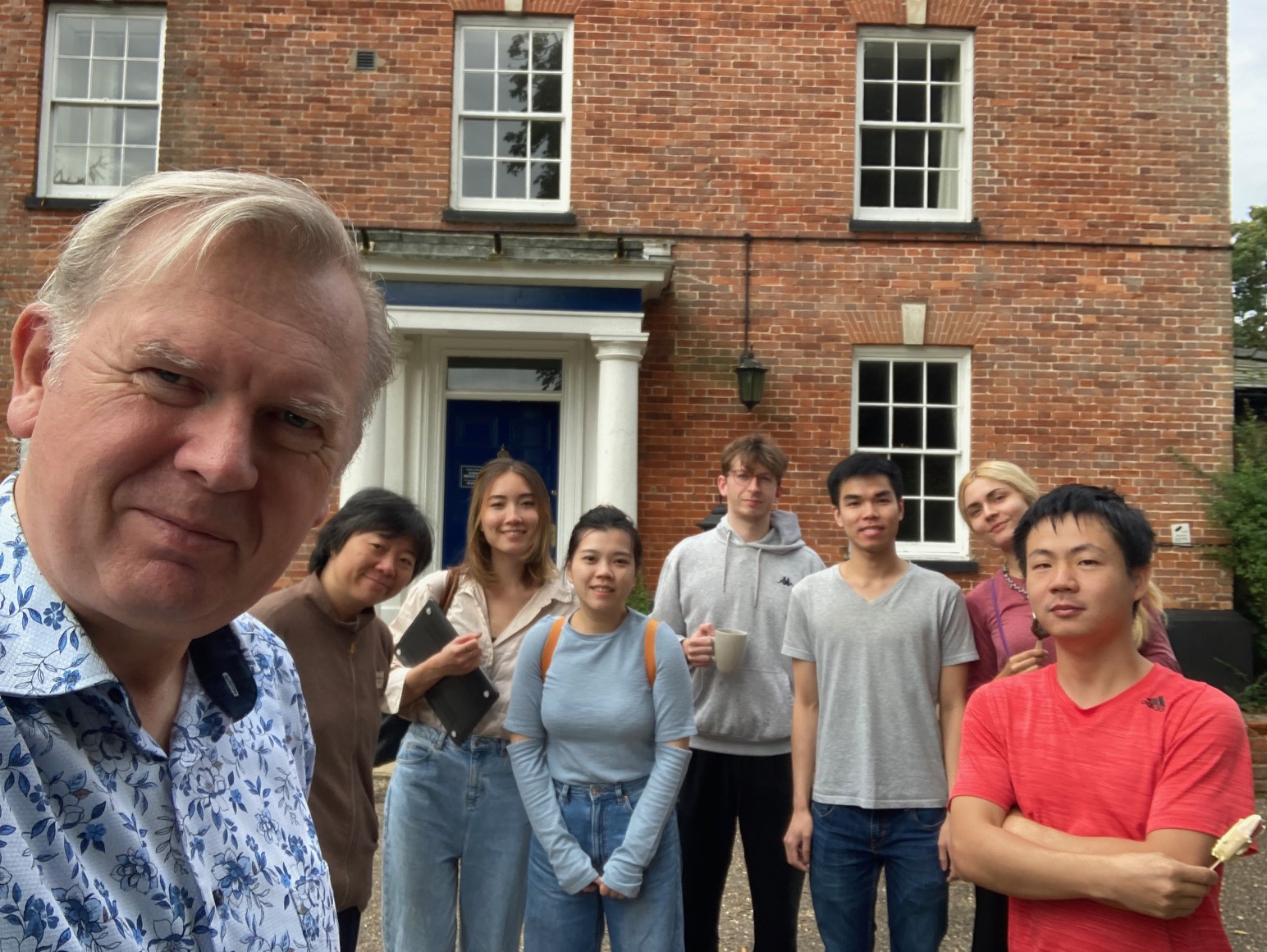
Labretreat to Swaffham, Norfolk. We had excellent brainstorms and have seen lots of seals during the hike! -
September 2023
AlphaFold Multimer predicts cross-kingdom interactions at the plant-pathogen interface. Excellent collaboration between Felix and Jie! -
September 2023
Dr. Issy! Issy submitted her thesis and passed her viva with only minor corrections. Matthieu Joosten was one of the examiners. Congrats, Issy! -
August 2023
Dr. Alice! Alice submitted her DPhil thesis and passed her viva with only minor corrections. Gail Preston and Tolga Buzkurt were her examiners. Well done Alice!!! -
June 2023
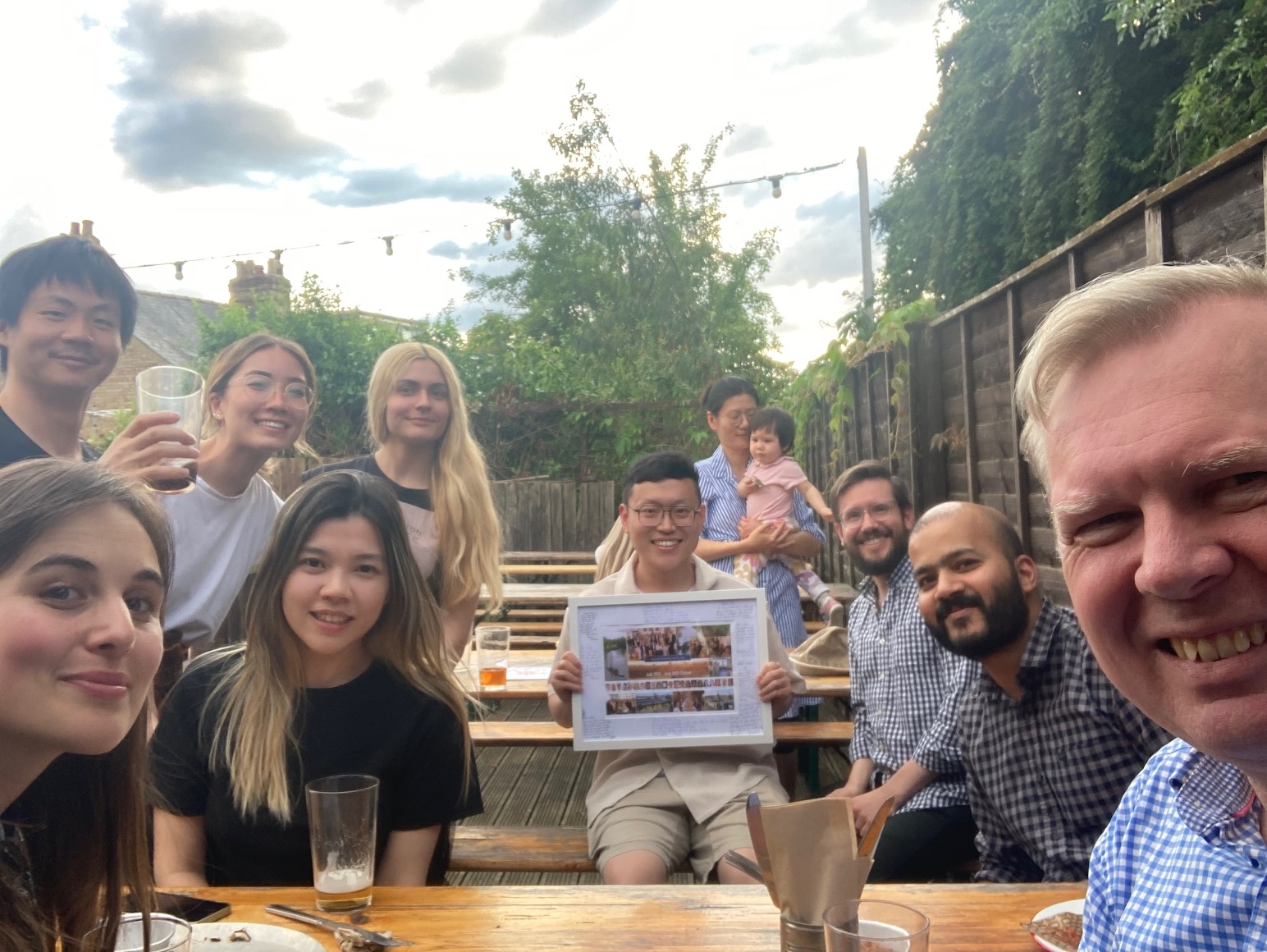
Bye Shijian! Only with us for 12 months, Shijian has been very productive on the molecular pharming project and returned to China to start a company! -
June 2023
Bye Y4 students! Sam, Ieuan, Alice and Edan were with use to do their Y4 projects with Shijian, Konstantina, Jie and Tee, respectively. They had a great time and made very interesting discoveries! -
May 2023
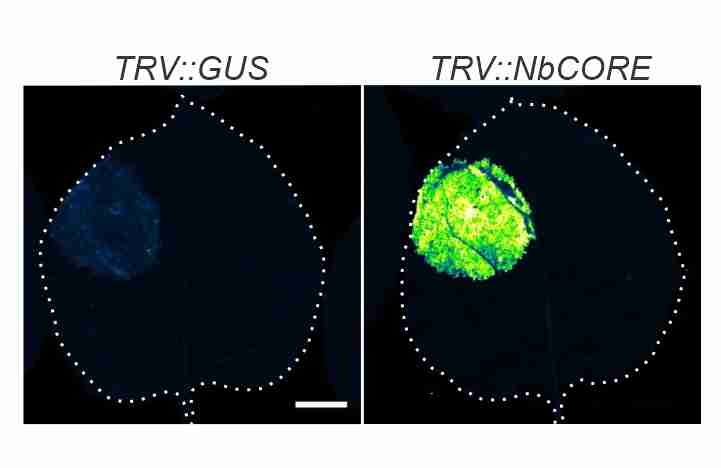
Issy's CORE paper! Depletion of CORE, the receptor of the cold shock protein of Agrobacterium results in increased transient protein expression! -
March 2023
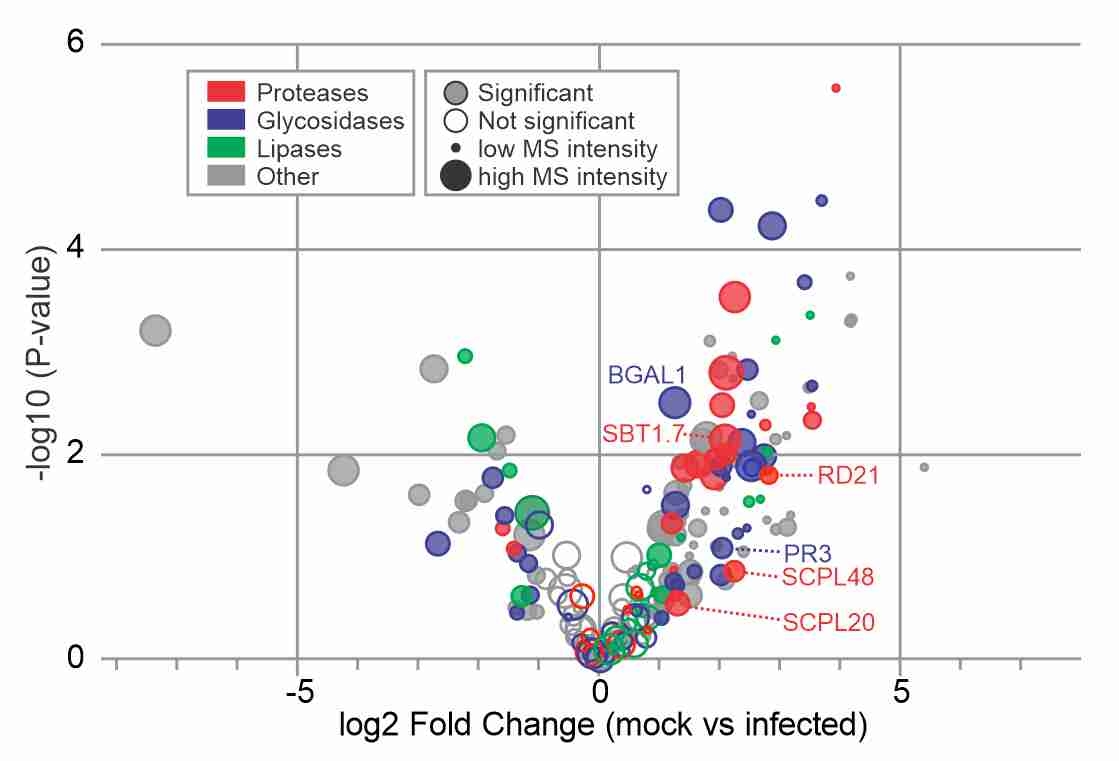
Suppressed hydrolases! Daniela and Alice published their manuscript in New Phytologist reporting dozends of secreted hydrolases that are suppressed during infection, and showing that one of them, PR3, is a neo-functionalized hydrolase that contributes to immunity.